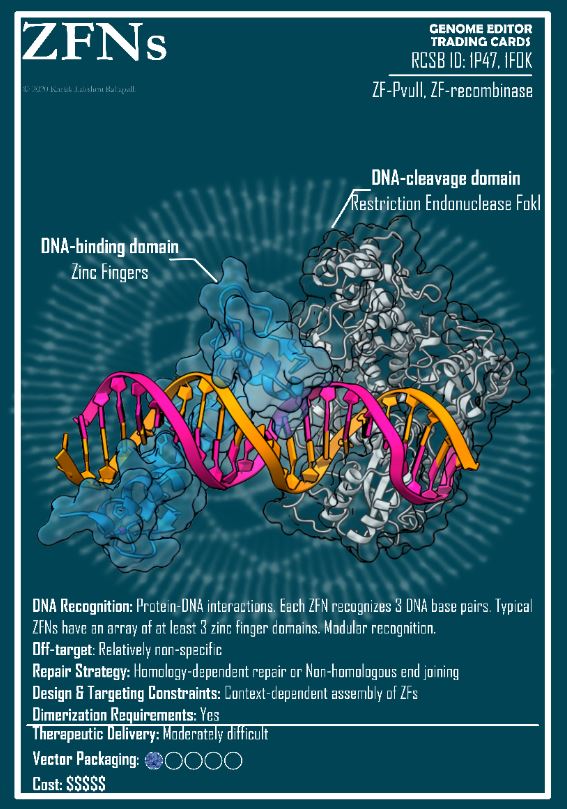
Modular design is the latest trend for developing new CRISPR tools. In The CRISPR Journal, Juan Carlos Collantes et al. present a base-editor system called Pin-point that recruits a DNA base-modifying enzyme through a hook (an RNA aptamer) within the guide-RNA molecule. In Nature Communications the goal of Lacramioara Bintu and colleagues is not base editing but epigenomic editing, the effector is a chromatin regulator and the hook is an antibody. When the CRISPR-effector combo is big, delivery of individual modules is easier. Furthermore, if the effector is already present inside the cell it can be simply recruited by providing the right hook. One more potential advantage is the convenient reconfiguration of the system by the mix and match of individual components and simultaneous recruitment of different effectors to different target sites.