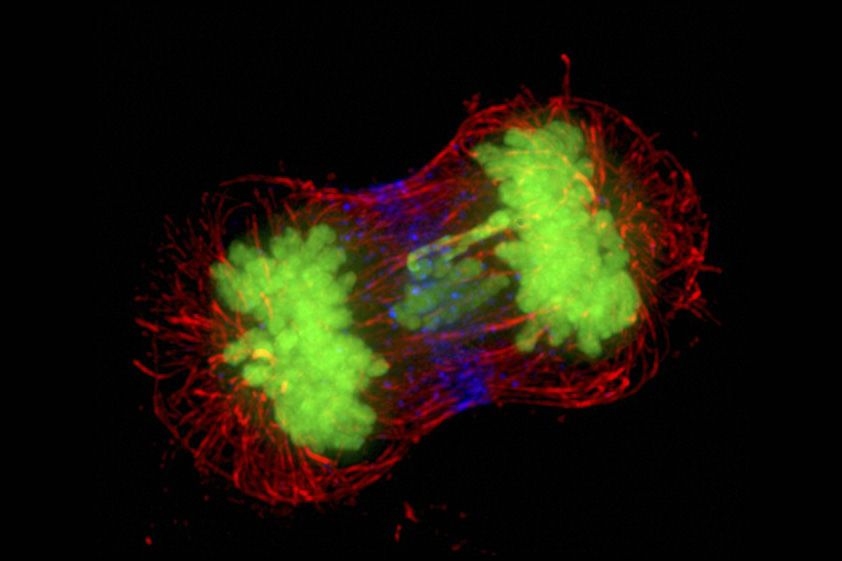
Humans have 23 pairs of chromosomes. The record-holder among animals (a butterfly called Polyommatus atlantica) can boast 229. Some plants have even more, but only because their genomes have undergone multiple rounds of duplication. We’re talking about chromosomes, of course. Their number is characteristic of each species and still shrouded in mystery. Why that number? And what would happen if we changed it?
In animals, the effects tend to be detrimental: mice with fused chromosomes, for instance, show abnormalities in behavior, growth, and fertility. Plants, however, appear surprisingly flexible, as demonstrated by a new experiment using CRISPR, recently published in Science.