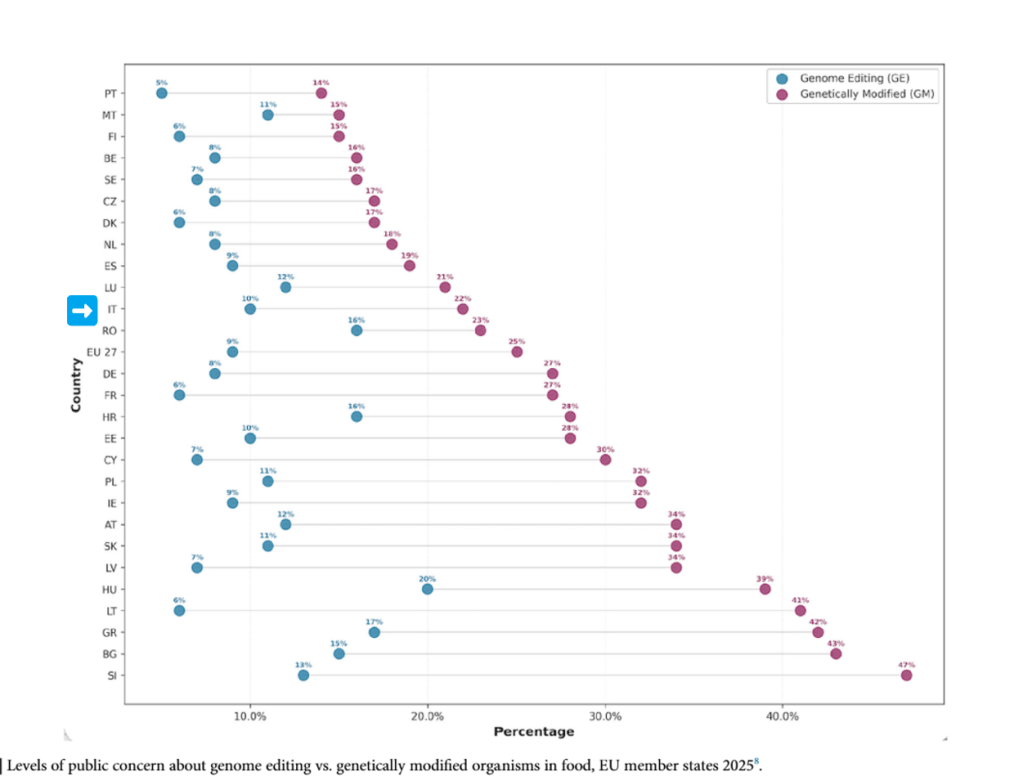
It won’t be long now: around the middle of the month we will find out whether the new European regulation on new genomic techniques (NGTs) will be approved in its current form or whether further compromises will be needed. This makes it a good moment to assess public perceptions of these technologies in agri-food applications. Are Europeans more open to NGTs than to traditional GMOs? The answers can be found in a survey commissioned by the European Commission at the request of the European Food Safety Authority (Eurobarometer 2025). The topic has returned to the spotlight thanks to an article published a few days ago by a group of experts on public perceptions of biotechnology in a journal of the Nature group devoted to sustainable agriculture.
Continue reading