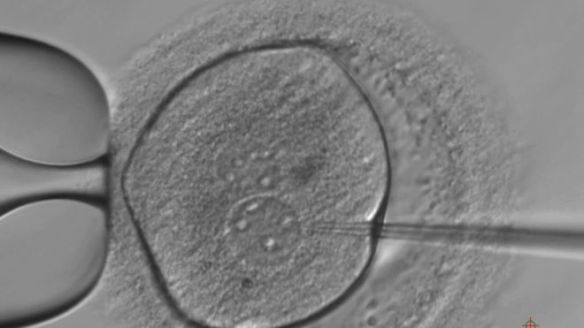
 They are the first human embryos edited in Europe and reported in scientific literature. The key difference with experiments already carried out in China and US is that the research published by Nature last week doesn’t have embryonic gene therapy in view. The London Francis Crick’s Institute team, in fact, was not interested in correcting disease-causing mutations but in increasing knowledge on human embryonic development. We asked one of the authors, Alessandro Bertero, to explain goals and results. The Italian researcher was pursuing his Ph.D. at Cambridge when he helped to refine the technique used by Kathy Niakan and colleagues to edit the genome of embryos. He answered our questions via Skype from America, where he continues working on embryonic stem cells as a postdoctoral fellow at Washington University
They are the first human embryos edited in Europe and reported in scientific literature. The key difference with experiments already carried out in China and US is that the research published by Nature last week doesn’t have embryonic gene therapy in view. The London Francis Crick’s Institute team, in fact, was not interested in correcting disease-causing mutations but in increasing knowledge on human embryonic development. We asked one of the authors, Alessandro Bertero, to explain goals and results. The Italian researcher was pursuing his Ph.D. at Cambridge when he helped to refine the technique used by Kathy Niakan and colleagues to edit the genome of embryos. He answered our questions via Skype from America, where he continues working on embryonic stem cells as a postdoctoral fellow at Washington University
What is the aim of the British approach to human embryo editing?
Embryo survival rate is low after in vitro fertilization, 10% in the third month of pregnancy. Many women must undergo 4 or 5 cycles, which are physically and psychologically heavy, as well as expensive. Hopefully, the IVF success rate could be boosted by improving the way embryos are cultured, increasing some growth factor or decreasing others. But finding the right formula requires a better understanding of how key genes affect embryonic development in the human species. Culture procedures developed so far are grounded mainly on mouse experiments and subsequent empirical adjustments. Probably this is one of the reasons why some pregnancies fail.
Why did you target the OCT4 gene?
There are three reasons for this choice. Because this gene has the right timing expression, because of its phenotypic and functional effects, and finally because slight differences are expected between human and mouse embryos. The differences have proved to be considerable indeed. By blocking the expression of OCT4, development stops earlier in humans. If OCT4 is not working well, embryos at the blastocyst stage are like balloons which swell and deflate until collapsing. Perhaps some cells are not firmly connected, or some important signals may be missing. Molecular mechanisms have yet to be clarified.
Chinese researchers led the way by editing discarded embryos that were unfit for implantation, while the US team specifically created their embryos for research. What about yours?
A total of 58 supernumerary embryos were donated by couples who no longer need them for assisted reproduction. Regulatory oversight is stricter in the UK than in the US. The Human Fertilization and Embryology Authority carefully assessed the ethical issues. One could also argue ex-post that this kind of research on human embryos is ethical precisely because the mouse is not a perfect model for human embryo development.
What did you exactly do?
It was a great chance for me to work with researchers from the Crick Institute, they are real pioneers. I optimized the RNA guide used by the CRISPR system to locate the target gene and then cut it. Broken ends are then fixed by natural repairing mechanisms in the cell, introducing point mutations and stopping the gene expression as a result. I experimented on pluripotent human stem cells, then the procedure was tested on mouse embryos and finally on human embryos. This preparatory work lasted over a year.
The American experiment last August apparently marked a huge advancement in CRISPR efficiency and accuracy, but in the meantime the interpretation of results has been questioned. Did CRISPR work well in your embryos?
We used existing embryos so we could not inject CRISPR and the sperm at the same time as they did. Therefore, we have higher rates of mosaicism, meaning that different cells in the same embryo contain different genetic sequences. Efficiency, however, was pretty good: 45% of the embryos had the desired mutation in both the copies of the gene, at least 70% in one. Mosaics are a problem for germ-line gene therapy, where the purpose is to correct a genetic disease in all the cells of an individual. However, in developmental biology studies, this phenomenon can be used to investigate what happens with different gene expression profiles. These data are a mine of information waiting to be analyzed.
(Photo credit: Kathy Niakan)
